RECENT ADVANCES IN COMPUTATIONAL PREDICTIONS OF NMR PARAMETERS FOR STRUCTURE ELUCIDATION OF CARBOHYDRATES: METHODS AND LIMITATIONS
N. D. Zelinsky Institute of Organic Chemistry, Russian Academy of Sciences, Moscow, Russia
Chemical Society Reviews, 2013, v. 42, pp. 8376-8415
DOI: 10.1039/C3CS60073D, PMID: 23887200
The review sections about empirical NMR simulation and molecular mechanics/dynamics were signifgicantly abdridged to fit the journal scope.
Click here for the original version (PDF): Simulation of NMR observables of carbohydrates.
All living systems are comprised of four fundamental classes of macromolecules - nucleaic acids, proteins, lipids, and carbohydrates (glycans). Glycans play a unique role of joining three principal hierarchical levels of the living world: 1) molecular level (pathogenic agent and vaccine recognition by the immune system; metabolic pathways involving saccharides that provide cells with energy; and energy accumulation via photosynthesis); 2) nanoscale level (cell membrane mechanics; structural support of biomolecules; and glycosylation of macromolecules); 3) microscale and macroscale levels (polymeric materials, such as cellulose, starch, glycogen, and biomass). NMR spectroscopy is the most powerful research approach for getting insight into solution structure and function of carbohydrates at all hierarchical levels, from monosaccharides to oligo- and polysaccharides. Recent progress in computational procedures opened a novel opportunity to reveal structural information available in the NMR spectra of saccharides and to advance our understanding of corresponding biochemical processes. The ability to predict the molecular geometry and NMR parameters is crucial for elucidation of carbohydrate structures. In the present paper, we review the major NMR spectrum simulation techniques in regard to chemical shift, coupling constant, relaxation rate and nuclear Overhauser effect prediction applied to the three levels of glycomics. Outstanding development in the related fields of genomics and proteomics has clearly shown that it is the advancement of research tools (automated spectrum analysis, structure elucidation, synthesis, sequencing and amplification) that drives grand challenges in modern science. Combination of NMR spectroscopy and computational analysis of structural information encoded in the NMR spectra reveals the way to automated elucidation of the structure of carbohydrates.
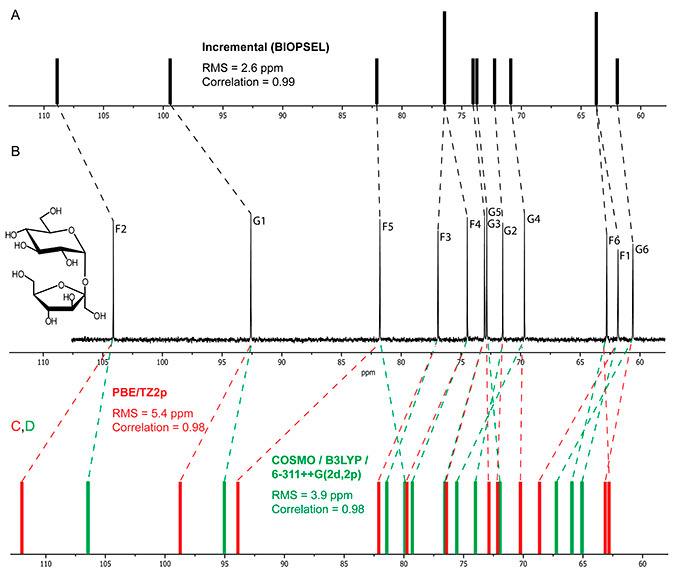
As a representative example, a figure from conlusion section demonstrates the accuracy of 13C chemical shift simulation of sucrose preformed at empirical and quantum mechanical levels: