BIOPSEL : structure generation
Basing on the monomeric composition, the program scans all the possible structures of the chemical repeating unit, excluding repetitions and considering the type of linkage that each residue may form. Monomeric residue structural properties are stored in easy-to-edit database, initially containing about 100 residues.
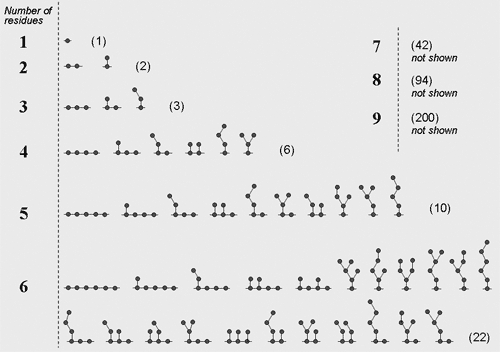
The first step and the outmost cycle of structures generation is working out all the possible topologies of the repeating unit (topolgies for mono- to hexameric units are presented on the figure). The maximal number of residues per repeating unit is nine. If the partial structure is known, this limitation may be overcome by combining two or more residues into one and describing this one as a separate residue with certain structural and spectral properties. Some of rarely-occurring topologies may be excluded from calculation by the special program key (widespread mode) to improve the performance and to simplify the result interpretation.
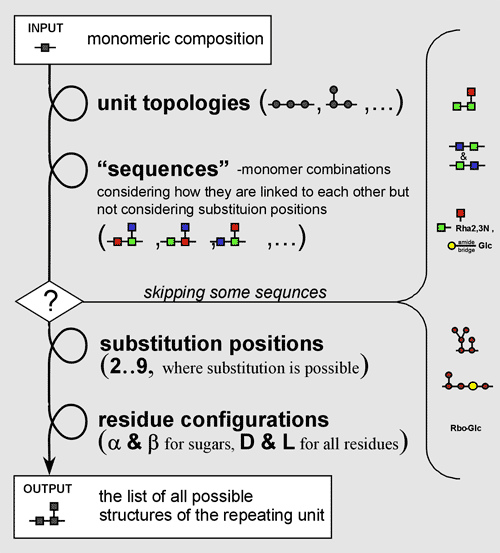
The second step is generation of so called sequences for each topology. The sequence is the monomer combination considering the order of how they are connected each to other, but ignoring the substitution positions and anomeric and absolute configurations. Some of sequences obtained may have no chemically-possible substitution pattern and, thereby, are be skipped.
|
The third step is generation of all chemically-possible substitution patterns for each sequence, considering the type of linkage that each substituting residue forms at C1. After this, anomeric and absolute configurations that have not been specified are iterated for each residue with unknown configuration.
The total number of generated structures of the prepeating unit depends on how many and what type of substitutable positions each residue has. The typical values are shown in the table:
| residues in the repeating unit |
unknown configurations |
simplifications + known substitution positions |
analyzed structures |
possible sequences, < |
calculation time, (Athlon 1700+) (hrs :) min : sec |
|---|---|---|---|---|---|
| 3 or less | 6 | 0 + 0 | < 44.0 K | < 14 | < 1 sec |
| 4 4 |
4 8 |
0 + 0 0 + 0 |
294.0 K 4.59 M |
108 108 |
Ā0 : 04 Ā1 : 11 |
| 5 5 4+Lys 5 5, widespread 5, widespread |
5 5 4 5 5 10 (=all) |
0 + 0 2 + 0 2 (except Lys) + 0 2 + 1 2 + 0 2 + 0 |
10.22 M 5.66 M 541.1 K 648.0 K 2.36 M 75.38 M |
1056 1056 1056 1056 384 384 |
Ā3 : 33 Ā2 : 03 Ā0 : 09 Ā0 : 02 Ā0 : 48 25 : 40 |
| 6 6 6, widespread 6, widespread 6, widespread |
0 5 5 5 5 |
1 + 0 3 + 0 3 + 0 3 + 1 3 + 5 |
21.91 M 379.77 M 92.63 M 21.15 M 102.0 K |
13.13 K 13.13 K 2640 2640 2640 |
Ā9 : 50 2 : 44 : 48 38 : 20 10 : 36 Ā0 : 02 |
| 7 7, widespread 7, widespread |
7 7 7 |
4 + 3 4 + 3 4 + 0 |
159.57 M |
191.3 K 20.39 K 20.39 K |
~15 hrs * 1 : 10 : 56 ~50 hrs * |
| 7+Lys 7+Lys, widespr. 8, widespread |
7 7 0 |
4 (except Lys) + 3 4 (except Lys) + 3 4 + 0 |
102.30 M |
3.25 M 182.1 K 182.1 K |
~32 hrs * 52 : 12 ~27 hrs * |
| 8+P 8+P, widespread 8+P, widespread |
4 8 8 |
5 (except P) + 8 5 (except P) + 8 5 (except P) + 5 |
25.22 M 108.28 M |
64.22 M 1.77 M 1.77 M |
48 : 25 1 : 01 : 53 ~120 hrs * |
The widespread mode forces the program to analyze only widespread sequences, skipping rarely-occurring ones.
The simplification is the inability of residue to be substituted at a certain position as compared to hexapyranose, e.g. a FucpNAc residue introduces two simplifications as it can not be substituted at C2 and C6, and Ribo residue does not introduce any simplifications as it can not be substituted at C6 but can be substituted at C5. Any simplifications fasten the calculation, as well as presence of phosphate groups, identical residues, alditols and amide bonds, especially in widespread mode.
Last update: 2002