UPDATES TO THE SYMBOL NOMENCLATURE FOR GLYCANS (SNFG) GUIDELINES
|
1State University of New York at Buffalo, USA 2Soka University, Japan 3National Library of Medicine (NLM), USA 4Biognos AB Göteborg, Sweden 5Swiss Institute of Bioinformatics (SIB), Switzerland 6GIP GmbH, Offenbach, Germany 7NextMove Software, USA 8Macquarie University, Australia 9Albert Einstein College of Medicine, USA 10Zelinsky Institute of Organic Chemistry, Russia 11University of California, San Diego, USA 12University of Georgia, Athens, GA, USA |
13Other members of the SNFG Discussion group: Alan Darvill, University of Georgia, USA Anne Dell, Imperial College London, UK Bernard Henrissat, Centre National de la Recherche Scientifique (CNRS), France Carolyn Bertozzi, Stanford University, USA Gerald Hart, University of Georgia, USA Hisashi Narimatsu, Research Center of Medical Glycoscience, Japan Hudson Freeze, Sanford-Burnham-Prebys Research Institute, USA Issaku Yamada, The Noguchi Institute, Japan James Paulson, Scripps Research Institute, USA James Prestegard, University of Georgia, USA Jamey Marth, , University of California Santa Barbara, USA JFG Vliegenthart, Bijvoet Center, Netherlands Marilynn Etzler, UC Davis, USA Markus Aebi, ETH Zürich, Switzerland | Minoru Kanehisa, Kyoto University, Japan Naoyuki Taniguchi, Riken Global Research Cluster, Japan Nathan Edwards, Georgetown University, USA Pauline Rudd, National Institute for Bioprocessing Research & Training, UK Peter Seeberger, Max-Planck-Institute of Colloids and Interfaces, Germany Raja Mazumder, The George Washington University, USA Rene Ranzinger, University of Georgia, USA Richard Cummings, Harvard Medical School, USA Ronald Schnaar, Johns Hopkins University School of Medicine, USA Serge Perez, French National Centre for Scientific Research, France Stuart Kornfeld, Washington University in St. Louis, USA Taroh Kinoshita, Osaka University, Japan William York, University of Georgia, USA Yuriy Knirel, Russian Academy of Sciences, Russia |
KEYWORDS: database, glycoscience, glycobiology, software, symbol nomenclature
Glycobiology, 2019, т. 29(9), стр.620-624
DOI: 10.1093/glycob/cwz045, PMID: 31184695
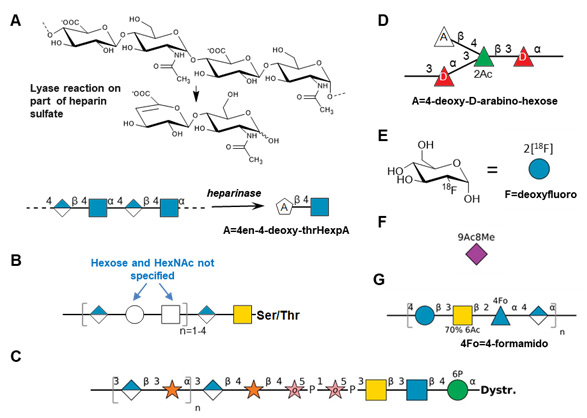
The Symbol Nomenclature For Glycans (SNFG) is a community-curated standard for the depiction of monosaccharides and complex glycans using various colored-coded, geometric shapes, along with defined text additions. It is hosted by the National Center for Biotechnology Information at the NCBI-Glycans Page (www.ncbi.nlm.nih.gov/glycans/snfg.html). Several changes have been made to the SNFG page in the last year to update the rules for depicting glycans using the SNFG, to include more examples of use, particularly for non-mammalian organisms, and to provide guidelines for the depiction of ambiguous glycan structures. This Glycoforum article summarizes these recent changes.